| Martinos Center: |
| Program in Cardiovascular Magnetic Resonance |
| Home |
| Mission |
| Research Initiatives |
| People |
| Facilities |
| Links |
| Available Positions |
| Contact |
 |
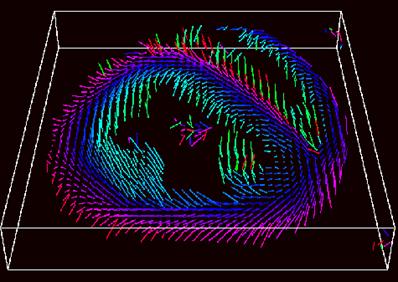
Our group has had a long interest in the development of novel magnetic resonance techniques to characterize myocardial structure and mechanics at the microscopic level. These techniques can be divided in to those designed to image myocardial fiber architecture and ultrastructure, and those designed to perform true 3-D measurements of myocardial strain. The fiber architecture of the myocardium is highly ordered but also highly complex and anisotropic. It is now well documented, for instance, that the orientation of myofibrils with respect to the long axis of the heart spans a range of approximately 1200 from endocardium to epicardium. In addition, a growing body of evidence suggests that the loss of normal fiber architecture may be one of the earliest signs of myocardial damage. We have therefore developed the ability to perform Diffusion Tensor Imaging (DTI) of the myocardium in-vivo, where the eigenvectors and eigenvalues of the tensor indicate the principal axes of diffusion within the myocardium. Since diffusion is far greater along myocardial fibers than through them, this data can then be used to construct maps of in-vivo myocardial fiber architecture. The principal technical challenge in acquiring this data, however, is the need for a technique that can separate spin dephasing due to diffusion from the dephasing due to the motion of the heart during the cardiac cycle. We have used a double-gated stimulated echo sequence, with an EPI readout, to overcome this challenge, and have successfully been able to map myocardial fiber architecture in ex-vivo specimens as well as in patients in-vivo.
In addition to myocardial diffusion tensor imaging, our group has also developed a technique to calculate myocardial strain rate from a 3 x 3 tensor of velocity differentials. Strain is inherently three- dimensional, and a 3-D measurement of strain is also coordinate invariant. The ability to acquire true 3D datasets for strain rate tensor imaging has been facilitated by the development of a technique by our group that allows for the simultaneous acquisition of two adjacent 2-D slices of velocity encoded MR data. The two adjacent slices can be combined to form a 3D dataset from which velocity differentials and hence strain rate can be calculated. This 2 slice method allows the calculation of each 3D strain rate tensor component from a single time coherent data set. This greatly increases strain rate sensitivity and significantly improves the quality of the strain rate image data as compared to conventional multi-slice methods.
Several different schemes have been used to display the
data derived from the strain rate tensor. For
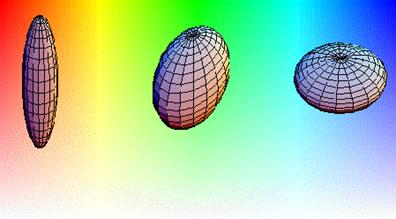
instance a hue, saturation, luminescence (HSL) color scheme, shown below, has been
developed to allow the strain eigenvalues associated
with the principal eignenvectors to be interpreted in
a physiologically meaningful manner. This color scheme maps the character of shape change to hue, the degree of change to saturation and relevant anatomy to luminescence. The change of a myofibril in systole from
a relaxed elongated state to a contracted shortened state is described by
negative strain along one of the principal eigenvectors and by positive strain
along the other two, while diastole is described by the reverse values.
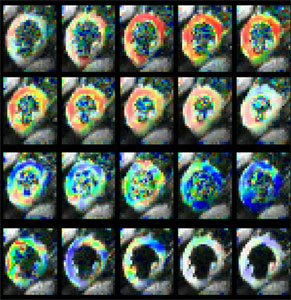
Strain rate tensors can be constructed several times in a single heartbeat, by using a rapid EPI readout, thus allowing myocardial strain rate to be followed over the cardiac cycle. The concatenation of strain rate images acquired with varying trigger delays allows myocardial strain rate to be imaged with a high temporal resolution from datasets needing only a few breath-hold scans to be acquired. It should be noted that the strain rate data acquired with this technique, in addition to being truly three dimensional, also have a significantly higher spatial resolution than data acquired with tagging techniques.
The novel techniques described above have the potential to provide new insights into the pathogenesis of myocardial dysfunction, and facilitate the diagnosis of myocardial damage at a far earlier stage. Clinical trials utilizing these techniques are currently being conducted in collaboration with the cardiology division at the
|