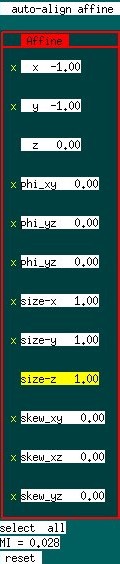
An affine transformation maintains all parallel lines in the original volume as parallel lines in the transformed volume,
http://en.wikipedia.org/wiki/Affine_transformation
and includes rigid-body translation (3 parameters) and rotation (3) plus uniform dilation/contraction (3) and skews (3), for a total of 12 parameters. The algorithm begins by using large parameters steps ("level 3") and proceeds to two lower levels with finer steps. Adjustment begins with a rigid-body motion and then incorporates other parameters.
Parameters sets
Parameters that will be adjusted automatically include those with an "x" by the name. By default, all parameters are adjusted. To include or exclude a parameter interactively, use up/down arrows to go the parameter, and then hit "x" to toggle the inclusion of the parameter on/off. Note that the parameter set can be specified at startup time from the unix/linux command line. Refer to startup options.
Also, parameter values can be initialized at startup from the unix/linux command line, which is a useful way to incorporate multi-step alignment into scripts. E.g., perform an initial alignment in a rodent using an anatomical scan, and then perform a second alignment step on distorted EPI data (with matching slices) by starting with the first parameter set but eliminating the slice position (z) and scaling (size-z) from the parameter set, because the slice-selection step is the same for anatomical and EPI data.
Increments
Increments on parameter values are automatically defined according to the template resolution. To adjust increments for a given parameter, highlight the parameter and hit the "tab" key. Then the left-right arrows will adjust parameter increments rather than parameter values.
Initialization
It's generally required to achieve a rough beginning alignment - perhaps within 1/4 of the field of view - before automated alignment. After adjusting the basic orientation on the setup page, one can manually adjust affine parameters as necessary by highlighting a parameter and using left/right arrows to modify the values, which are reflected in the updated image.